-
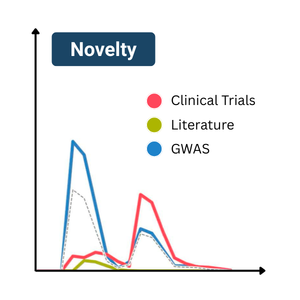
A metric for target novelty
December 2025
Published in Nature Communications, an Open Targets team developed a metric for the novelty of a target in the context of a disease, according to current available knowledge. This allows drug discovery scientists to easily identify potentially novel targets. It also reveals a shift in how genetic evidence is used in drug discovery.
-
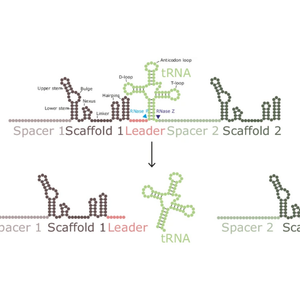
A next-generation dual guide CRISPR system for genetic interaction library screening
December 2025
Publicshed in Nature Communications, a team at Open Targets and the Wellcome Sanger Institute have created a next-generation system for dual guide delivery, enabling efficient library-scale screening for hundreds of thousands of genetic interactions.
-
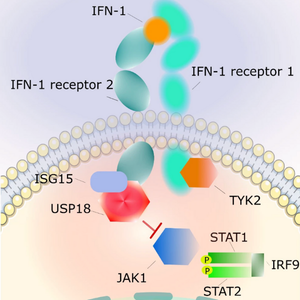
Largest trans-eQTL meta-analysis in a single cell type prioritises USP18 in lupus
October 2025
Published in Nature Communications, this is the largest trans-eQTL meta-analysis in a single cell type. An Open Targets team analysed 3,734 lymphoblastoid cell line samples across nine cohorts, identifying four robust loci.
-
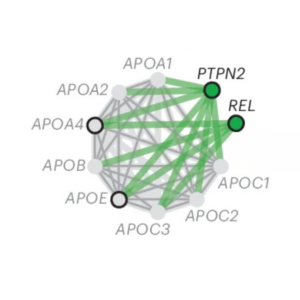
A tissue-specific atlas of protein–protein associations enables prioritisation of candidate disease genes
May 2025
Published in Nature Biotechnology, a new in silico method to predict protein associations based on co-abundance creates an atlas of protein associations across 11 human tissues
-
Cancer drug resistance causes and categories identified
October 2024
In a study published in Nature Genetics, researchers have mapped the genetic landscape of cancer drug resistance, uncovering that DNA changes can be grouped into four main categories and highlighting possible new therapeutic targets.
-
Why Clinical Trials Stop: the role of genetics
July 2024
Using machine learning to analyse the genetic factors behind early clinical trial termination, researchers find a link between genetic evidence and trial outcome.
-
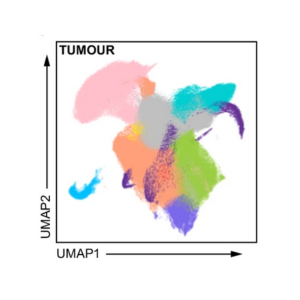
An immune atlas of lung cancer
May 2024
Published in Nature Communications, Open Targets researchers and collaborators used single-cell RNA sequencing to create a high resolution molecular map of immune cells in NSCLC tumours, to better understand the role of these cells in disease progression.
-
Researchers decipher mechanisms of liver regeneration
May 2024
Published in Nature, the team studied liver regeneration during chronic disease, and showed that chronic injury creates an environment that induces cellular plasticity in human organs, paving the way for regenerative therapies.
-

MSD joins Open Targets
April 2024
MSD, the tradename of Merck & Co., Inc., Rahway, N.J., USA, joins the Open Targets consortium.
-

Open Targets turns 10!
March 2024
Open Targets celebrates the 10th anniversary of its founding.
-
Cancer drug discovery accelerated as hundreds of overlooked targets prioritised
January 2024
The second generation of the Cancer Dependency Map uncovers 370 priority drug targets, with strong links to specific cancer types.
-

David Hulcoop appointed Open Targets Director
July 2023
Open Targets has appointed David Hulcoop as its new Executive Director, following the retirement of Ian Dunham. Hulcoop will build from the programme’s existing capabilities to maximise Open Targets impact on drug discovery decision making.
-
A human interactome to prioritise drug discovery
February 2023
In a study published in Nature Genetics, researchers created a network of interacting proteins with which they identified groups of proteins interacting with genes that have been linked through GWAS to over 1,000 human traits from 21 therapeutic areas.
-
Bowel cancer mutations that impact immunotherapy identified
January 2023
Open Targets researchers use CRISPR and mini tumours to discover more about how mutations in the IFN-gamma immune pathway affect the development of colorectal cancer.
-

Genentech joins Open Targets
November 2022
Genentech, a subsidiary of the Roche Group, brings its expertise in biotechnology to the consortium.
-

Single-cell mapping of T cells create unique map of activation process
May 2022
Continuing their previous work published in 2019, a team of Open Targets researchers create a one-of-a-kind, detailed map of gene expression dynamics during the T cell activation process, uncovering new biology and prioritising targets for new therapies.
-
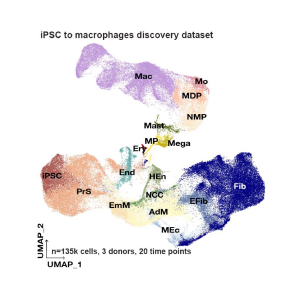

Roadmap of myeloid differentiation will help better understand immunity
May 2022
Scientists at Open Targets create a molecular map of myeloid differentiation using over 470,000 human induced pluripotent stem cells, showing that the differentiated cells are able to acquire definitive tissue-resident identities.. This will be an important resource to explore myelopoiesis and new therapeutic opportunities.
-

Pfizer joins Open Targets
February 2022
Pfizer will contribute its unique expertise in oncology, immunology, and metabolic disorders, complementing the expertise of our five current partners.
-
Defining the PROTACtable genome
July 2021
An Open Targets project, published in Nature Reviews Drug Discovery, established a new framework to assess whether human proteins could be targeted with Proteolysis Targeting Chimeras (PROTACs).
-

Mapping how gene expression varies in human microglia
June 2021
Open Targets researchers define how factors such as age, sex, and clinical pathology affect microglia, a critically important cell in the development and diseases of the human central nervous system.
-

Next-Generation Platform released
April 2021
Our new, next-generation Open Targets Platform is the culmination of over two years of work. It reinvents the best of the old Platform with a redesigned experience, improved usability, based on powerful modern technologies.
-
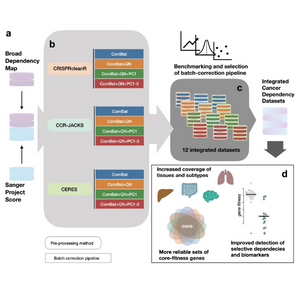
Largest integrated resource of pan-cancer CRISPR-Cas9 screens to date
March 2021
For the first time, a collaboration between Open Targets and the Broad Institute integrated independent CRISPR-Cas9 screens performed at each institution, providing greater statistical power to cancer- and subtype-specific analyses.
-
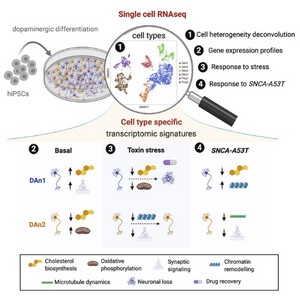
A systematic survey of dopaminergic neuron differentiation biases
March 2021
A systematic survey of 215 iPS cell lines obtained from HipSci identified molecular signatures to select the best lines to use in the study of neuronal development, unlocking the potential of iPSC lines to study otherwise inaccessible cell states.
-

Getting a finer view of genes leading to Alzheimer's disease risk
February 2021
An Open Targets paper identifies 37 regions associated with Alzheimer’s disease, including 4 novel ones, and prioritises a list of genes within those regions that may alter Alzheimer’s disease risk.
-
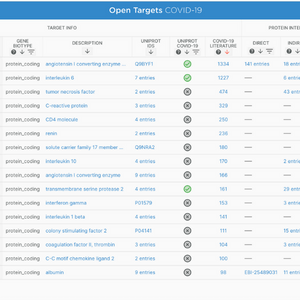
COVID-19 Prioritisation tool launched
July 2020
In response to the COVID-19 pandemic, the Open Targets team created a tool to support drug target identification and drug repurposing for COVID-19. The information contained in this platform has since been integrated into the Open Targets Platform.
-
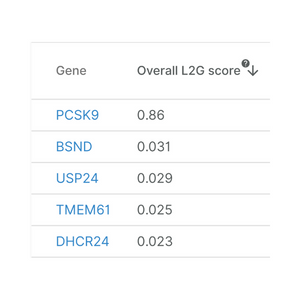
Locus-to-gene method introduced
April 2020
The Locus-to-gene (L2G) machine-learning scoring method is integrated in the Open Targets Genetics Platform. It reflects the likelihood that a gene is causal for the trait in question. The method was published in Nature Genetics in October 2021.
-

eQTL Catalogue launched
January 2020
Open Targets researchers created a database of uniformly processed gene expression and splicing QTLs from a wide range of human studies. The method was published in September 2021.
-
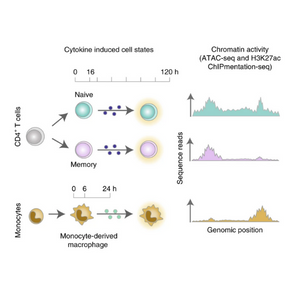
Analysis of chromatin activity identifies T cell states driving complex immune diseases
September 2019
A team of researchers at Open Targets analyse the chromatin activity of immune disease-associated variants at different stages of T cell and macrophage activation, and show that such variants have a role in early rather than late activation of memory T cells.
-

Open Targets celebrates 5-year anniversary
June 2019
Colleagues from all of our consortium partners gathered at the Wellcome Genome Campus in Hinxton, UK to celebrate 5 years of Open Targets and plan the milestones for our next 5 years.
-
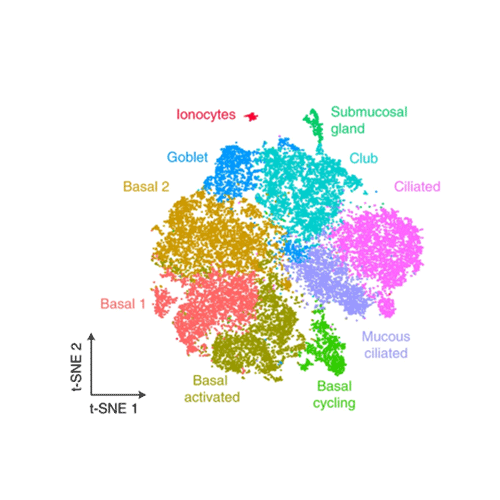
Single cell lung map published in Nature Medicine
June 2019
Open Targets researchers map health and asthmatic lung tissue to uncover new drug targets for treating asthma
-
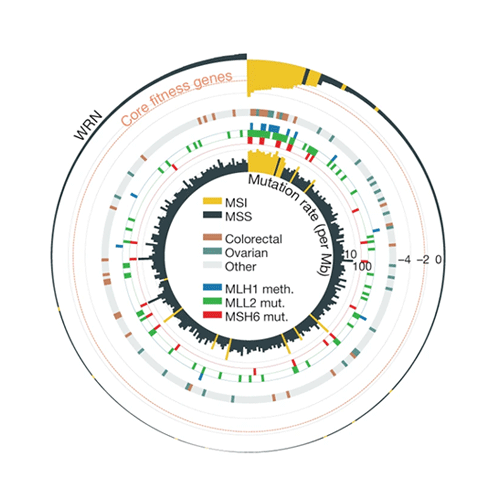
Large-scale release of CRISPR data from Project Score publication
April 2019
Open Targets publishes results from its landmark experimental project where CRISPR technology was used in over 300 cancer models to discover thousands of key genes essential for cancer’s survival.
-

Ian Dunham appointed Open Targets Director
January 2019
Ian Dunham is appointed Director for Open Targets and will continue to focus on the delivery of our established research programme that aims to exploit advances in genetics and genomics for drug target identification and prioritisation.
-

Sanofi joins Open Targets
October 2018
Sanofi joins the consortium and brings its expertise in immunology, oncology, neurosciences and diabetes to help us identify potential drug targets.
-
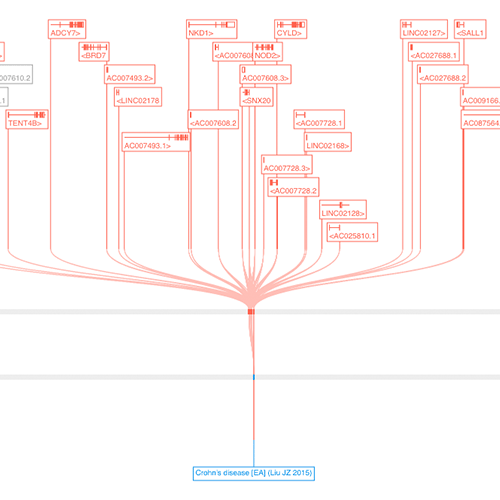
Open Targets Genetics release
October 2018
During ASHG 2018, we officially released Open Targets Genetics, a new web tool that allows users to browse gene-variant-trait relationships from UK Biobank data and GWAS Catalog studies.
-

Rolf Apweiler appointed Interim Director
August 2018
Rolf Apweiler becomes the Interim Director for Open Targets and continues to foster new partnerships and collaborations to drive drug discovery.
-

Celgene joins Open Targets
May 2018
Celgene joins Open Targets and brings its expertise in developing innovative therapies for patients with cancer, immune-inflammatory, and other unmet medical needs.
-

Takeda joins Open Targets
December 2017
Takeda joins Open Targets and brings its expertise in oncology, gastroenterology and central nervous system therapeutic areas, along with vaccines.
-

New brand, new data: introducing Open Targets
April 2016
To celebrate the release of our first experimental data, CTTV rebrands itself as Open Targets but maintains its focus on helping researchers identify and prioritise potential therapeutic drug targets.
-

Biogen joins the Centre for Therapeutic Target Validation (CTTV)
February 2016
Biogen joins Open Targets and brings its expertise in innovative therapies for the treatment of neurodegenerative diseases, hematologic conditions and autoimmune disorders.
-
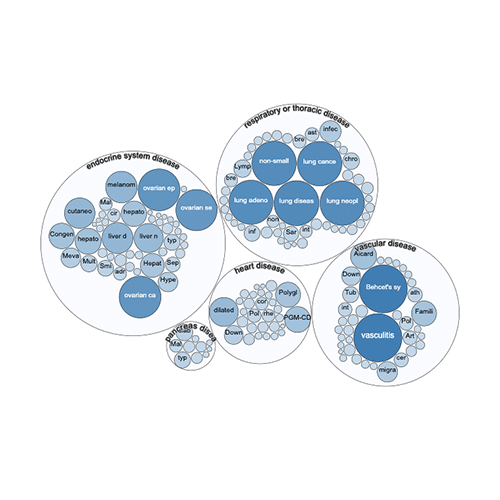
Target Validation Platform launched
December 2015
Underscoring our commitment to share our data openly to benefit the broader scientific community, we launched a new web platform to help researchers identify therapeutic targets for new and repurposed medicines.
-

Jeffrey Barrett appointed founding Director
February 2015
After leading the development of the Immunochip genotyping array and playing an important role in UK10K Dr Jeffrey Barrett is appointed as the founding Director of the Centre for Therapeutic Target Validation (CTTV)
-

Centre for Therapeutic Target Validation launched
March 2014
The predecessor to Open Targets was founded in March 2014 when the European Bioinformatics Institute (EMBL-EBI), the Wellcome Sanger Institute, and GSK came together to build a partnership that would focus on target identification, prioritisation, and validation.